Vectors
|
VH, VK and VL primers
Competent cells
|
Yeast display vectors
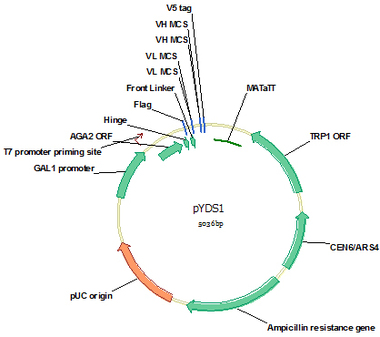
pYDS1 (Cat#: mAb001)
pYDS1 is a yeast display vector for monomer scFv display on yeast strain EBY100 surface. AGA1 is yeast cell surface protein and AGA2 is coupled to AGA1 via 2 disulfide bounds. Via fusion to C terminal of AGA2 through a hinge, scFv is displayed on yeast surface. A flag tag at the N terminus and a V5 tag at the C terminus of scFv were designed for convenient detection of scFv expression and antigen binding by flow cytometry. The (G4S)3 flexible linker was designed as the intralinker between VL and VH. This vector was designed for stepwise clone of VL and VH and all the enzymes sites for VH and VL cloning do not appear in VH and VL germline genes and the translated amino acids are favorable for forming a flexible linker.
pYDS1 is a yeast display vector for monomer scFv display on yeast strain EBY100 surface. AGA1 is yeast cell surface protein and AGA2 is coupled to AGA1 via 2 disulfide bounds. Via fusion to C terminal of AGA2 through a hinge, scFv is displayed on yeast surface. A flag tag at the N terminus and a V5 tag at the C terminus of scFv were designed for convenient detection of scFv expression and antigen binding by flow cytometry. The (G4S)3 flexible linker was designed as the intralinker between VL and VH. This vector was designed for stepwise clone of VL and VH and all the enzymes sites for VH and VL cloning do not appear in VH and VL germline genes and the translated amino acids are favorable for forming a flexible linker.
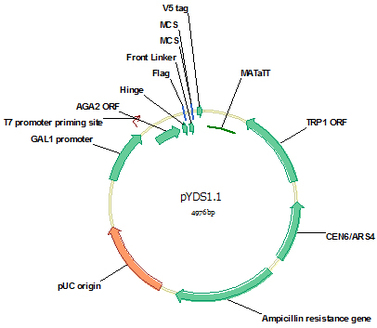
pYDS1.1 (Cat#: mAb002)
pYDS1.1 is similar to pYDS1 for monomer scFv display except that the multiple clone sites for VL and VH are shortened to 9 bp, which allows direct purification of the linearized vector using PCR purification kit without the necessity to remove the 9 bp insert by gel separation. This vector was designed for one step clone of both VL and VH either as assembled scFv or through three fragment gap repair. Yeast has a highly efficient homologous recombination system, which will efficiently assemble the co-transformed linearized vector, VL and VH into correct vector-scFv form through the homologous ends designed for the three fragments.
pYDS1.1 is similar to pYDS1 for monomer scFv display except that the multiple clone sites for VL and VH are shortened to 9 bp, which allows direct purification of the linearized vector using PCR purification kit without the necessity to remove the 9 bp insert by gel separation. This vector was designed for one step clone of both VL and VH either as assembled scFv or through three fragment gap repair. Yeast has a highly efficient homologous recombination system, which will efficiently assemble the co-transformed linearized vector, VL and VH into correct vector-scFv form through the homologous ends designed for the three fragments.
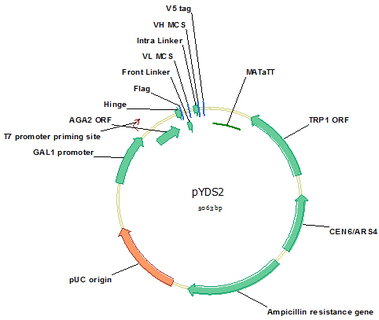
pYDS2 (Cat#: mAb003)
If the eventual goal of the isolated scFv is to be engineered to dimmers, such as scFv-Fc or full antibody, the dimmer-scFv library is recommended because high affinity monomer scFv may not translate to higher affinity dimmers. Direct isolation of scFvs that have high affinities in dimmer form is a wise approach. pYDS2 is designed for this need. Through the natural 3 disulfide bounds in the hinge region, dimmer scFvs are displayed on yeast surface. pYDS2 was designed for stepwise cloning of VL and VH like pYDS1. All the tags, linkers and enzyme sites are the same as pYDS1 for efficient gene clone, flexible linkers, and convenient scFv detection by flow cytometry.
If the eventual goal of the isolated scFv is to be engineered to dimmers, such as scFv-Fc or full antibody, the dimmer-scFv library is recommended because high affinity monomer scFv may not translate to higher affinity dimmers. Direct isolation of scFvs that have high affinities in dimmer form is a wise approach. pYDS2 is designed for this need. Through the natural 3 disulfide bounds in the hinge region, dimmer scFvs are displayed on yeast surface. pYDS2 was designed for stepwise cloning of VL and VH like pYDS1. All the tags, linkers and enzyme sites are the same as pYDS1 for efficient gene clone, flexible linkers, and convenient scFv detection by flow cytometry.
Yeast secretory expression vectors
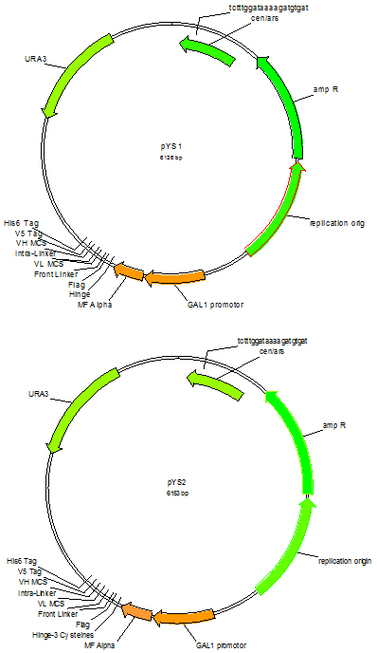
pYS1 (Cat#: mAb004) and pYS2 (Cat#: mAb005)
After enrichment of antigen specific yeast display populations through magnetic and flow sortings, high affinity individual scFc clones can be identified using flow cytometry. However, this is time and labor consuming and the display scFvs need to be converted to soluble form later for affinity and other characterizations. It would be more efficient if converting the display scFv to soluble scFv then use high throughput ELISA to identify individual clones. Vectors pYS1and pYS2, counterparts for pYDS1, pYDS1.1 and pYDS2, were designed for soluble scFv expression. Simply amplify scFv gene from the enriched display yeast population using PCR and co-transform into YVH10 yeast strain with linearized vector pYS1, pYS1.1 or pYS2, pick individual clones, culture and induce in 96 deep well plates, the scFv containing supernatants are ready for ELISA identification. The positive clones could be expanded from the 96 well stock for large scale scFv purification for further characterization. Vector pYS1 pairs with pYDS1 and pYDS1.1 for monomer scFv expression, and pYS2 pairs with pYDS2 for dimmer scFv expression.
After enrichment of antigen specific yeast display populations through magnetic and flow sortings, high affinity individual scFc clones can be identified using flow cytometry. However, this is time and labor consuming and the display scFvs need to be converted to soluble form later for affinity and other characterizations. It would be more efficient if converting the display scFv to soluble scFv then use high throughput ELISA to identify individual clones. Vectors pYS1and pYS2, counterparts for pYDS1, pYDS1.1 and pYDS2, were designed for soluble scFv expression. Simply amplify scFv gene from the enriched display yeast population using PCR and co-transform into YVH10 yeast strain with linearized vector pYS1, pYS1.1 or pYS2, pick individual clones, culture and induce in 96 deep well plates, the scFv containing supernatants are ready for ELISA identification. The positive clones could be expanded from the 96 well stock for large scale scFv purification for further characterization. Vector pYS1 pairs with pYDS1 and pYDS1.1 for monomer scFv expression, and pYS2 pairs with pYDS2 for dimmer scFv expression.
Phage display vectors
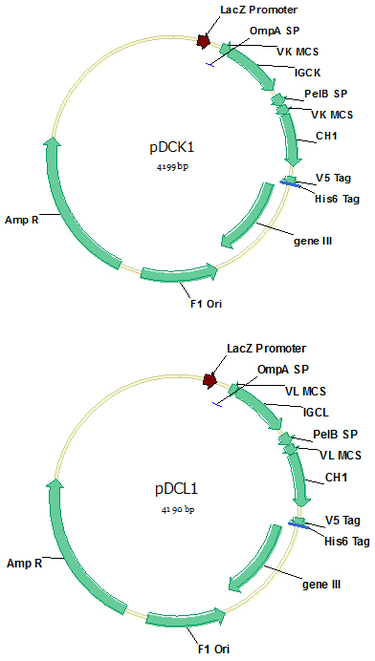
pDCK1 (Cat#: mAb006) and pDCL1 (Cat#: mAb007)
Vectors pDCK1 and pDCL1 contain a CH1 constant region of IgG1, as well as a kappa and a lambda constant region respectively, for Fab display simply by cloning VL/VK and VH into the vectors. All the constant regions were codon-optimized for E.coli expression, and all the enzymes sites designed for efficient digestion and ligation do not appear in antibody V genes. Long stuff sequences have been designed for the VK/VL and VH clone enzymes for efficient enzyme digestion. These two vectors could also be used for phage display scFv library construction using Xba I / Not I digestion.
Vectors pDCK1 and pDCL1 contain a CH1 constant region of IgG1, as well as a kappa and a lambda constant region respectively, for Fab display simply by cloning VL/VK and VH into the vectors. All the constant regions were codon-optimized for E.coli expression, and all the enzymes sites designed for efficient digestion and ligation do not appear in antibody V genes. Long stuff sequences have been designed for the VK/VL and VH clone enzymes for efficient enzyme digestion. These two vectors could also be used for phage display scFv library construction using Xba I / Not I digestion.
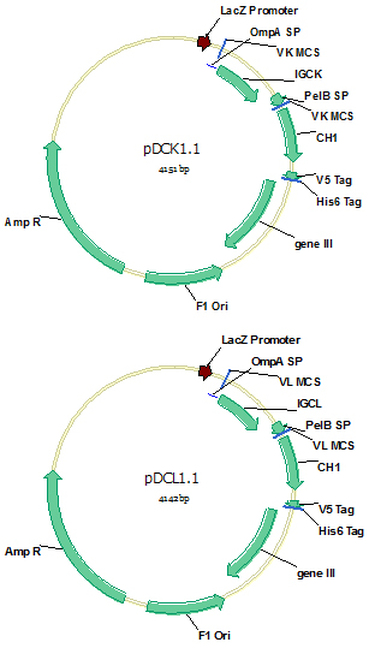
pDCK1.1 (Cat#: mAb008) and pDCL1.1 (Cat#: mAb009)
Vectors pDCK1.1 and pDCL1.1 were the same as pDCK1 and pDCL1 respectively except that the stuff sequence for the VK/VL and VH clone enzymes were shortened to 9bp so that digested vector could be conveniently purified by PCR purification kit instead of the labor consuming gel separation.
Vectors pDCK1.1 and pDCL1.1 were the same as pDCK1 and pDCL1 respectively except that the stuff sequence for the VK/VL and VH clone enzymes were shortened to 9bp so that digested vector could be conveniently purified by PCR purification kit instead of the labor consuming gel separation.
Bi-cistronic lentiviral vectors
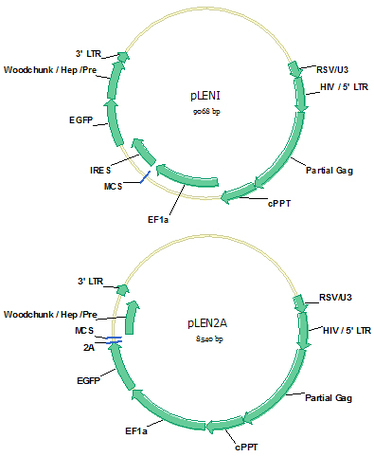
pLENI (Cat#: mAb010) and pLEN2A (Cat#: mAb011)
Establishment of antigen positive cell lines is necessary for cell based antibody library panning and the validation of isolated antibody afterwards. Lentiviral vectors pLENI and pLEN2A are designed for bicistronic expression of protein of interest and EGPF simultaneously. EGFP will be expressed intracellularly for convenient observation of successful viral infection and gene expression, as well as to purify high expression cells by FACS (Fluorescence Activated Cell Sorting). If the protein of interest is cell membrane protein, the translated protein will locate on cell surface. Vector pLENI uses a IRES (Internal ribosome entry site) sequence to link the gene of interest and EGFP gene to ensure the same mRNA expression level of the two proteins; while vector pLEN2A uses a 2A sequence to ensure the same protein expression level of the two proteins, which will be separated by enzyme cleavage in 2A site in endosome. Lentivirus could be easily made by co-transduction of pLENI or pLEN2A with vectors providing VSVG, REV and Gag/pol protein expression into HEK293 cells.
Establishment of antigen positive cell lines is necessary for cell based antibody library panning and the validation of isolated antibody afterwards. Lentiviral vectors pLENI and pLEN2A are designed for bicistronic expression of protein of interest and EGPF simultaneously. EGFP will be expressed intracellularly for convenient observation of successful viral infection and gene expression, as well as to purify high expression cells by FACS (Fluorescence Activated Cell Sorting). If the protein of interest is cell membrane protein, the translated protein will locate on cell surface. Vector pLENI uses a IRES (Internal ribosome entry site) sequence to link the gene of interest and EGFP gene to ensure the same mRNA expression level of the two proteins; while vector pLEN2A uses a 2A sequence to ensure the same protein expression level of the two proteins, which will be separated by enzyme cleavage in 2A site in endosome. Lentivirus could be easily made by co-transduction of pLENI or pLEN2A with vectors providing VSVG, REV and Gag/pol protein expression into HEK293 cells.
E.coli expression vector
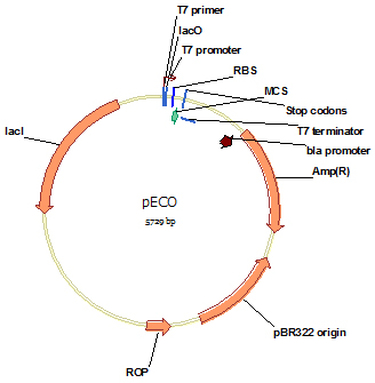
pECO (Cat#: mAb012)
Vector pECO is designed for high-level intracellular expression of a gene of interest by E.coli from the strong bacteriophage T7 promoter. All the enzymes are designed for convenient and efficient gene clone. There is no tag designed for this vector. For convenient purification, a His6 tag could be designed at N or C end of the protein of interest depend on the need. Protein expression is induced by IPTG.
Vector pECO is designed for high-level intracellular expression of a gene of interest by E.coli from the strong bacteriophage T7 promoter. All the enzymes are designed for convenient and efficient gene clone. There is no tag designed for this vector. For convenient purification, a His6 tag could be designed at N or C end of the protein of interest depend on the need. Protein expression is induced by IPTG.
Mammalian cell expression vectors
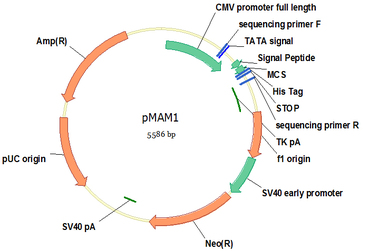
pMAM1 (Cat#: mAb013)
Vector pMAM1 is designed for highly efficient secretory protein expression in mammalian cells, such as CHO or HEK293 cells. Protein expression is driven by the potent CMV promoter, led by a VH3 signal peptide that is native for most antibody secretion in B lymphocytes , and followed by a His8 tag for convenient protein purification. Enzymes in multiple clone site are designed for convenient and efficient gene clone. Recombinant protein could be directly purified from the supernatant. If using serum free mammalian cell expression system, one step Nickel resin purification gives rise to high purity proteins. A neo gene expression cassette allows stable expression cell line establishment by G418 selection.
Vector pMAM1 is designed for highly efficient secretory protein expression in mammalian cells, such as CHO or HEK293 cells. Protein expression is driven by the potent CMV promoter, led by a VH3 signal peptide that is native for most antibody secretion in B lymphocytes , and followed by a His8 tag for convenient protein purification. Enzymes in multiple clone site are designed for convenient and efficient gene clone. Recombinant protein could be directly purified from the supernatant. If using serum free mammalian cell expression system, one step Nickel resin purification gives rise to high purity proteins. A neo gene expression cassette allows stable expression cell line establishment by G418 selection.
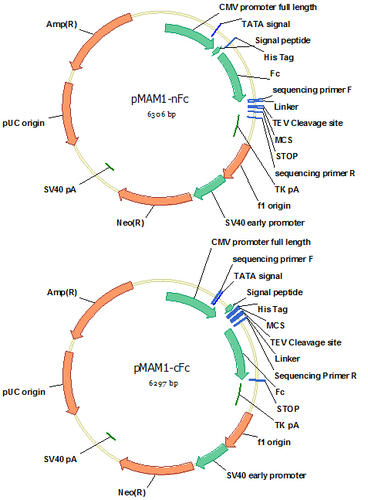
pMAM1-nFc (Cat#: mAb014) and pMAM1-cFc (Cat#: mAb015)
Serum free mammalian expression system allows convenient and efficient purification of high purity proteins if they are secreted outside of the cells. It is usually easy to express proteins to the supernatant if they are native secretory proteins. However, in many cases, intracellular proteins or partial protein, such as the extracellular domain of a cell surface protein, are difficult to be expressed in the secretory pathway, most likely due to the change of their native translocation mechanisms. Fusion of the difficult-expression proteins to an easy expression protein, such as Fc, is a solution because an easy-expression protein can work as a carrier to bring the difficult-expression protein out of the cells. Vectors pMAM1-nFc and pMAM1-cFc are designed for this purpose. They are modified based on pMAM1 by introduction of a human IgG1 Fc fragment at N or C terminus of the protein of interest. For convenient purification, a His8 tag is designed at the N terminus of the fusion protein. To avoid possible conformational hindrance between Fc and the protein of interest, a flexible (G4S)3 linker is designed between Fc and the multiple clone site (MCS). Fc fragment can also be removed from the fusion protein by TEV cleavage at the site designed between the two proteins. Preparation of the extracellular domain of a cell surface antigen is often very challenging. These two fusion vectors are capable of high level expression of extracellular domains of most cell surface proteins.
Serum free mammalian expression system allows convenient and efficient purification of high purity proteins if they are secreted outside of the cells. It is usually easy to express proteins to the supernatant if they are native secretory proteins. However, in many cases, intracellular proteins or partial protein, such as the extracellular domain of a cell surface protein, are difficult to be expressed in the secretory pathway, most likely due to the change of their native translocation mechanisms. Fusion of the difficult-expression proteins to an easy expression protein, such as Fc, is a solution because an easy-expression protein can work as a carrier to bring the difficult-expression protein out of the cells. Vectors pMAM1-nFc and pMAM1-cFc are designed for this purpose. They are modified based on pMAM1 by introduction of a human IgG1 Fc fragment at N or C terminus of the protein of interest. For convenient purification, a His8 tag is designed at the N terminus of the fusion protein. To avoid possible conformational hindrance between Fc and the protein of interest, a flexible (G4S)3 linker is designed between Fc and the multiple clone site (MCS). Fc fragment can also be removed from the fusion protein by TEV cleavage at the site designed between the two proteins. Preparation of the extracellular domain of a cell surface antigen is often very challenging. These two fusion vectors are capable of high level expression of extracellular domains of most cell surface proteins.
Human VH, VL and VK primers (Cat#: mAb016)
Based on the largest antibody database IMGT, we have designed a set of degenerative primers that are capable of amplifying all the human antibody repertoires. The forward primers anneal to the first frame region of V gene and reverse primers anneal to J regions. Totally, 7 forward and 3 reverse primers are designed for VH; 5 forward and 4 reverse primers are designed for VK and 9 forward and 5 reverse primers are designed for VL. A reverse primer for IgM is also available for naive library construction.
E.coli Competent cells
TOP10 (Cat#: mAb017) (20 reactions for $200, Chemical)
TOP10 Chemically Competent E. coli are provided at a transformation efficiency of 1 x 109 cfu/µg plasmid DNA in individual tubes. They are ideal for high-efficiency cloning and plasmid propagation. They allow stable replication of high-copy number plasmids.
BL21 (Cat#: mAb018) (20 reactions for $200, Chemical)
BL21 Chemically Competent cells are provided at a transformation efficiency of 1 x 108 cfu/µg plasmid DNA in individual tubes. BL21 E. coli cells are ideal for high-level expression of non-toxic recombinant proteins.
TG1 (Cat#: mAb019) (20 reactions for $200, Electroporation)
TG1 is the standard cell for phage display library construction. With a high efficiency of 1 × 1010 cfu/µg plasmid DNA, our TG1 electrocompetent cells are perfect for phage display antibody or peptide library construction.
TOP10 Chemically Competent E. coli are provided at a transformation efficiency of 1 x 109 cfu/µg plasmid DNA in individual tubes. They are ideal for high-efficiency cloning and plasmid propagation. They allow stable replication of high-copy number plasmids.
BL21 (Cat#: mAb018) (20 reactions for $200, Chemical)
BL21 Chemically Competent cells are provided at a transformation efficiency of 1 x 108 cfu/µg plasmid DNA in individual tubes. BL21 E. coli cells are ideal for high-level expression of non-toxic recombinant proteins.
TG1 (Cat#: mAb019) (20 reactions for $200, Electroporation)
TG1 is the standard cell for phage display library construction. With a high efficiency of 1 × 1010 cfu/µg plasmid DNA, our TG1 electrocompetent cells are perfect for phage display antibody or peptide library construction.
Loading buffer
DNA loading buffer (10 x) (Cat#: mAb020) (3 x 1 ml, $30)
DNA Loading Buffer (10X) is a liquid buffer for easy loading and tracking of DNA samples in
agarose or native polyacrylamide gels.
Protein loading buffer (6x) (Cat#: mAb021) (10 ml, $15)
Laemmli is a sample buffer to use in protein electrophoresis and western blot. Every one volume 6X sample buffer is added to five volumes of protein sample and is boiled or heated for 5-10 minutes.
DNA Loading Buffer (10X) is a liquid buffer for easy loading and tracking of DNA samples in
agarose or native polyacrylamide gels.
Protein loading buffer (6x) (Cat#: mAb021) (10 ml, $15)
Laemmli is a sample buffer to use in protein electrophoresis and western blot. Every one volume 6X sample buffer is added to five volumes of protein sample and is boiled or heated for 5-10 minutes.
